Key Points
-
The mitochondrial unfolded protein response (UPRmt) is a conserved transcriptional response activated by multiple forms of mitochondrial dysfunction and regulated by mitochondrial-to-nuclear communication.
-
The UPRmt promotes cell survival and mitochondrial recovery through metabolic adaptations and a precise mitochondrial biogenesis programme.
-
UPRmt activation promotes lifespan extension and protects against bacterial pathogens that perturb mitochondrial function.
-
Prolonged UPRmt activation can lead to the propagation of deleterious mitochondrial genomes and mitochondrial damage and can contribute to age-associated organismal deterioration.
-
UPRmt manipulations are potential therapeutic targets for treating the vast number of diseases associated with mitochondrial dysfunction.
Abstract
Mitochondrial function declines during ageing owing to the accumulation of deleterious mitochondrial genomes and damage resulting from the localized generation of reactive oxygen species, both of which are often exacerbated in diseases such as Parkinson disease. Cells have several mechanisms to assess mitochondrial function and activate a transcriptional response known as the mitochondrial unfolded protein response (UPRmt) when mitochondrial integrity and function are impaired. The UPRmt promotes cell survival and the recovery of the mitochondrial network to ensure optimal cellular function. Recent insights into the regulation, mechanisms and functions of the UPRmt have uncovered important and complex links to ageing and ageing-associated diseases. In this Review, we discuss the signal transduction mechanisms that regulate the UPRmt and the physiological consequences of its activation that affect cellular and organismal health during ageing.
Similar content being viewed by others
Main
Mitochondria are membrane-bound organelles that are derived from the engulfment of an α-proteobacterium by a pre-eukaryotic cell more than 1 billion years ago1,2. Like their bacterial ancestor, mitochondria have an outer membrane and an inner membrane that separates the intermembrane space (IMS) from the matrix. Mitochondria harbour small circular genomes (mtDNA) that encode 13 oxidative phosphorylation (OXPHOS) proteins as well as tRNAs and ribosomal RNAs required for their synthesis, which occurs in the mitochondrial matrix3. The remainder of the ∼1,200 mitochondrial proteins are encoded by nuclear genes, synthesized on cytosolic ribosomes and subsequently targeted to and imported into mitochondria4.
Mitochondria are best known for being the cellular powerhouses that generate the majority of cellular energy by coupling nutrient oxidation to ATP synthesis via the respiratory chain and ATP synthase. In addition to energy production, mitochondria have other essential metabolic roles, including in amino acid and nucleotide synthesis, iron–sulfur cluster biogenesis, intermediate metabolite biogenesis and calcium homeostasis. Furthermore, mitochondria can function as signalling platforms to regulate programmed cell death and innate immunity5.
Mitochondrial and OXPHOS dysfunction is the primary cause of a group of heterogenous disorders known as mitochondrial diseases that most commonly impair neuron or muscle cell function and are typically caused by mutations in genes that encode OXPHOS proteins or genes required for the assembly of the respiratory chain and ATP synthase complexes6. Mitochondrial function also declines during ageing owing to the accumulation of damaged mtDNA, increased reactive oxygen species (ROS) generation, toxin exposure and general organelle deterioration. Mitochondrial dysfunction is thought to contribute to the decline in cellular function observed during ageing and to loss of muscle strength and neuronal function7,8,9,10,11. Age-associated mitochondrial dysfunction is often exacerbated or accelerated in neurodegenerative diseases such as Parkinson disease and Alzheimer disease7,8.
The mitochondrial unfolded protein response (UPRmt) is a transcriptional response that is activated by multiple forms of mitochondrial dysfunction and regulated by mitochondrial-to-nuclear communication12,13,14,15,16,17,18,19,20. Cells use a variety of mechanisms to monitor the health of the intracellular mitochondrial network, which is comprised of constantly dividing and fusing organelles to facilitate compartment mixing21 and ensure localized mtDNA replication in each mitochondrion22. If mitochondrial function declines, the UPRmt is activated to promote the repair and recovery of this network and maintain cellular function13,15,18,23.
A nuanced relationship between ageing and the UPRmt has become increasingly apparent. For example, the UPRmt promotes repair and recovery from transient mitochondrial dysfunction caused by toxins or pathogens15,16,24 and by mutations in nuclear-encoded respiratory chain genes13,18. However, aberrant or prolonged UPRmt activation results in cells being in a state of constant mitochondrial recovery that can increase the accumulation of deleterious mitochondrial genomes harbouring loss-of-function point mutations or deletions (ΔmtDNAs) during animal development and ageing that could directly contribute to age-associated deterioration25,26. Thus, it is important that the UPRmt is precisely regulated. Recent work has revealed that the UPRmt is regulated at multiple levels, including at the levels of transcription and chromatin remodelling. Interestingly, chromatin remodelling during mitochondrial dysfunction enables more robust UPRmt activation in aged cells and promotes longevity27,28. In addition to intracellular UPRmt regulation, UPRmt signalling occurs between cells and tissues, presumably to regulate mitochondrial adaptations throughout an organism13,29,30,31.
In this Review, we discuss recent insights into the signalling mechanisms that regulate the UPRmt, including how the integrity of the mitochondrial network is assessed and the changes in transcription and chromatin organization that occur in order to maintain a functional mitochondrial network. We describe the effects of the UPRmt on cellular and organismal physiology that promote mitochondrial recovery and cell survival during mitochondrial dysfunction and the interactions with other mitochondrial stress responses such as mitophagy. We also discuss the relationship between age-associated mitochondrial deterioration in metazoans and the loss of UPRmt effectiveness with age13,32,33.
Activation and regulation of the UPRmt
The discovery of an adaptive stress response that functions to alleviate the accumulation of unfolded proteins within mitochondria was reported more than 15 years ago in cultured mammalian cells. Overexpression of an irreversibly misfolded mitochondrial protein resulted in increased accumulation of nuclear transcripts encoding several mitochondrial chaperones and proteases, presumably to maintain protein homeostasis (proteostasis) within mitochondria20. As this mitochondrial- specific response is conceptually similar to the transcriptional response activated by protein misfolding in the endoplasmic reticulum known as the UPR, it was named the UPRmt (Refs 19,34).
Over the past 10 years, genetic and biochemical studies, largely carried out using Caenorhabditis elegans and mammalian model systems, have elucidated the function of the UPRmt beyond the maintenance of mitochondrial proteostasis, as well as the mechanisms by which mitochondrial dysfunction is detected and communicated to the nucleus to regulate chromatin structure and transcriptional outputs.
A working definition of the UPRmt. Numerous studies have reported diverse cellular alterations associated with mitochondrial dysfunction. Here, we define the UPRmt as an ensemble of genes and cellular activities that are transcriptionally regulated by defined transcription factors that are activated following mitochondrial dysfunction. Mitochondrial dysfunction broadly refers to a wide range of defects that include the accumulation of unfolded proteins within the mitochondrial matrix but also conditions that perturb OXPHOS activity, such as genotoxic stress and chemical inhibitors. Below, we discuss recent insights supporting this definition, which continues to rapidly evolve as new discoveries are made.
Mitochondrial network assessment and nuclear communication in C. elegans. Using transcriptional reporters activated by mitochondrial dysfunction, multiple screens in C. elegans have identified factors that are required for UPRmt activation14,16,35,36,37,38. Consistent with a role in signalling between mitochondria and the nucleus, these factors are localized in mitochondria, the cytosol and the nucleus.
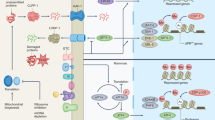
Activating transcription factor associated with stress (ATFS-1) is one of at least two transcription factors required for UPRmt activation in worms14. In addition to a nuclear localization sequence (NLS; common to all bZIP transcription factors), ATFS-1 has an amino-terminal mitochondrial-targeting sequence (MTS)18. This arrangement enables cells to evaluate the function of the mitochondrial network on the basis of mitochondrial protein import efficiency (Fig. 1). In cells with a healthy mitochondrial network, the MTS prevails, and ATFS-1 is imported into mitochondria, where it is degraded by the matrix-localized protease LON. When mitochondria are damaged, the NLS directs ATFS-1 to the nucleus to regulate transcription18. Importantly, MTS removal or inactivation results in constitutive nuclear accumulation of ATFS-1 and UPRmt activation18,39. Thus, compartmentalization of ATFS-1 regulates its transcriptional activity. Moreover, if ATFS-1 fails to be imported into mitochondria, the transcription factor becomes active, indicating that mitochondrial import is a key UPRmt regulatory event18.
Oxidative phosphorylation (OXPHOS) perturbation, excessive reactive oxygen species (ROS), impaired complex assembly (mitonuclear protein imbalance) and the accumulation of misfolded proteins impair mitochondrial protein import efficiency and activate ATFS-1 (activating transcription factor associated with stress). ATFS-1 is imported into healthy mitochondria via its mitochondrial-targeting sequence (MTS), where it is degraded. If mitochondrial import efficiency is perturbed, ATFS-1 is transported to the nucleus and the mitochondrial unfolded protein response (UPRmt) is activated. In the nucleus, ATFS-1 induces a number of genes that re-establish mitochondrial function and import efficiency, such as genes that promote recovery of the OXPHOS complexes, re-establish mitochondrial proteostasis by upregulating chaperones and proteases and detoxify ROS, as well as protein import components. NLS, nuclear localization sequence; TIM, translocase of the inner membrane; TOM, translocase of the outer membrane.
Most, if not all, conditions that activate the UPRmt are also known to influence mitochondrial protein import (Fig. 1; Table 1), which, as indicated by a growing body of evidence, occurs both post-translationally and co-translationally40. The majority of proteins targeted to the mitochondrial matrix have an amino-terminal MTS, which first targets them to the translocase of the outer membrane complex (TOM complex) on the mitochondrial outer membrane and then to the translocase of the inner membrane 23 complex (TIM23 complex) in the inner membrane41. Importantly, protein trafficking through the inner membrane and TIM23 complex requires ATP and the proton gradient generated by the respiratory chain, as well as the matrix-localized chaperone mtHSP70 (also known as HSPA9) and is impaired by high levels of ROS42,43,44. Various mitochondrial perturbations activate the UPRmt, including OXPHOS impairment by toxins16 or by RNAi-mediated knockdown of OXPHOS components and assembly factors13,45, impaired mitochondrial ribosome function following drug treatment or RNAi-mediated knockdown of genes encoding mitochondrial ribosomal proteins15 and depletion of mtDNA17,19 (Table 1). These perturbations induce mitonuclear protein imbalance, whereby the expression of stoichiometric mtDNA-encoded and nuclear-encoded OXPHOS components is mismatched, which results in the accumulation of OXPHOS components unable to assemble into complexes15. Unassembled OXPHOS components interact with mitochondrial chaperones and must be degraded by localized proteases to prevent toxic effects46,47. Similarly, impaired mitochondrial protein folding or inhibition of mitochondrial chaperones and proteases14,18,19 activates the UPRmt.
Owing to the pleiotropic nature of these mitochondrial defects, it is difficult to determine the precise events that lead to UPRmt activation, but they all impair mitochondrial protein import, consistent with the regulation of ATFS-1 by mitochondrial protein import (Fig. 1). These observations suggest that ATFS-1 is activated by mitochondrial dysfunction that ultimately feeds back on mitochondrial protein import efficiency owing to damage in the system. Consistent with this model, it was recently shown that in Saccharomyces cerevisiae, many mitochondrial proteins mislocalize to the cytosol when mitochondrial proteostasis and protein import are impaired. Interestingly, the mislocalized mitochondrial proteins are extremely toxic if they are not quickly degraded by cytosolic proteasomes48,49. These data suggest that mitochondrial import is impaired by multiple forms of mitochondrial dysfunction and that multiple pathways are activated in response to such dysfunction.
Mitochondrial network assessment and nuclear communication in mammals. Although the mechanisms of the UPRmt have been elucidated in C. elegans, the UPRmt was discovered in mammalian cells with the observation that mitochondrial chaperones and proteases are induced in response to mtDNA depletion or the accumulation of misfolded proteins in the mitochondrial matrix17,20. More recently, a UPRmt was found to be induced in mice and cultured mammalian cells by various mitochondrial perturbations, such as deletion of the mitochondrial aspartyl-tRNA synthetase DARS2 (Ref. 50), skeletal-muscle-specific deficiency in CRIF1, which is an integral protein of the large mitochondrial ribosomal subunit (39S)51, and inhibition of the mitochondrial ribosome, the mitochondrial protease LON and the mitochondrial chaperone tumour necrosis factor type 1 receptor-associated protein (TRAP1; also known as HSP75)15,52 (Table 1).
Although the resulting transcriptional response to mitochondrial dysfunction is similar in mammals and C. elegans, regulation is probably more complicated. Data obtained over the past 5 years suggest the involvement of three bZIP transcription factors, CHOP (also known as DDIT3), ATF4 and ATF5, associated with the integrated stress response (ISR)53,54,55,56. The expression of CHOP, ATF4 and ATF5 requires the phosphorylation of the eukaryotic translation initiation factor 2 subunit 1 (eIF2α), which is catalysed by four kinases responsive to diverse cellular stresses (Fig. 2). Some eIF2α molecules are constitutively phosphorylated, but the amount of phosphorylated eIF2α increases when the eIF2α kinases GCN2 and PERK (also known as EIF2AK3) are activated by amino acid depletion and endoplasmic reticulum dysfunction30,57,58,59, respectively. Increased eIF2α phosphorylation results in reduced global protein synthesis and preferential translation of mRNAs with open reading frames in the 5′ untranslated regions, such as those encoding CHOP, ATF4 and ATF5 (Refs 60,61,62).
Activation of the mammalian mitochondrial unfolded protein response (UPRmt) requires the phosphorylation of eukaryotic translation initiation factor 2 subunit 1 (eIF2α) by the eIF2α kinases GCN2 and PERK (also known as EIF2AK3), which are activated by amino acid depletion, excessive reactive oxygen species (ROS) or endoplasmic reticulum (ER) stress. Phosphorylation of eIF2α results in reduced protein synthesis and the preferential translation of mRNAs harbouring open reading frames in the 5′ untranslated region, such as the mRNAs encoding the transcription factors CHOP, ATF4 and ATF5. All three seem to be involved in the UPRmt, although it remains to be determined how their activity is regulated during mitochondrial dysfunction.
Multiple studies have demonstrated that diverse forms of mitochondrial stress induce the expression of CHOP, ATF4 and ATF5 as well as that of genes involved in the UPRmt, such as mitochondrial proteostasis and metabolic remodelling genes30,51,52,63,64,65,66. For example, mitochondrial chaperones and proteases, as well as CHOP, ATF4 and ATF5, were induced in mice deficient in DARS2 (Ref. 50). Multiple studies have also demonstrated that CHOP, ATF4 or ATF5 are required for the induction of UPRmt genes during mitochondrial dysfunction, consistent with a model in which these transcription factors regulate the mammalian UPRmt (Refs 30,51,64,66). Combined, these studies have rediscovered the ISR, but in the context of mitochondrial dysfunction. Almost 15 years ago, ATF4 and PERK were shown to mediate the transcriptional induction of mitochondrial proteostasis genes as well as genes involved in metabolic remodelling53. However, it is not yet clear how each ISR kinase and transcription factor is regulated during mitochondrial dysfunction to achieve a specific mitochondrial stress response, as expression of all three transcription factors can be activated during a variety of conditions, including endoplasmic reticulum stress, amino acid depletion and viral infection, which also increase eIF2α phosphorylation60,61,67,68,69,70,71.
A recent study suggested that ATF5 is a mammalian orthologue of ATFS-1, as ATF5 rescued UPRmt activation in worms lacking ATFS-1 (Ref. 64). This study also indicated that ATF5 can respond directly to mitochondrial stress because like ATFS-1, ATF5 activity seems to be regulated via mitochondrial import efficiency64. During mitochondrial stress, ATF5 was required for the induction of several mitochondrial chaperone and protease genes, consistent with an essential role of ATF5 in cell growth during mitochondrial stress64. Interestingly, ATF4 was also shown to protect against mitochondrial stress when cells were depleted of mtDNA72 or treated with either paraquat or a mitochondrial protease inhibitor66. Protection was mediated by induction of metabolic genes associated with the ISR3,66 (Table 2).
It is not clear how the functions of CHOP, ATF4 and ATF5 are coordinated during mitochondrial dysfunction. All three are bZIP proteins, but CHOP and ATF4 do not contain an MTS like ATFS-1 or ATF5 (Ref. 64), suggesting that they are regulated at the level of expression, which requires the induction of eIF2α phosphorylation56. One simple hypothesis is that CHOP, ATF4 and eIF2α phosphorylation are sufficient to induce ATF5 expression, consistent with studies indicating that transcription of ATF5 is dependent on CHOP and ATF4 (Refs 54,61,73,74). Once expressed, ATF5 activity can be modulated by mitochondrial function to regulate its transcriptional activity. However, ATF4 and CHOP probably have more direct roles as well.
When perturbations occur within the IMS, activation of the UPRmt requires the nuclear hormone receptor oestrogen receptor (ERα) in addition to CHOP, ATF4 and ATF5 (Refs 75,76). ERα activation leads to increased proteasome activity to eliminate mislocalized or defective IMS proteins. ERα-mediated signalling also increases expression of the IMS protease HTRA2 and the transcription factor nuclear respiratory factor 1 (NRF1), which promotes mitochondrial biogenesis77. Considering the variations in physiology between different mammalian tissues affected by mitochondrial dysfunction, it is possible that other mechanisms of mitochondrial-to-nuclear communication that activate a transcriptional response remain to be discovered.
Chromatin remodelling. Mitochondrial dysfunction stimulates chromatin remodelling required for UPRmt activation, which also requires mitochondrial-to- nuclear communication. Following mitochondrial stress, the cytosolic protein LIN-65 (which has unknown biochemical function) translocates to nuclei, which requires the cytosol-localized histone lysine methyltransferase MET-2 (homologous to the mammalian protein SETDB1). In the nucleus, LIN-65 and MET-2 promote chromatin compaction and global gene silencing during mitochondrial stress28. Concomitantly, the homeobox transcription factor DVE-1 (homologous to the mammalian DNA-binding proteins SATB1 and SATB2), which is also required for UPRmt activation35, specifically interacts with chromatin regions harbouring mitochondrial stress response genes to maintain a transcription-competent state and allow UPRmt activation28 (Fig. 3).
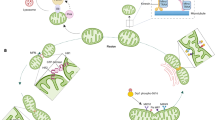
Mitochondrial stress leads to chromatin remodelling and global gene silencing. This requires LIN-65 and the incorporation of dimethylated histone H3 at Lys 9 (H3K9me2) in nucleosomes. Histone dimethylation is catalysed by the histone-lysine N-methyltransferase MET-2 in the cytosol . Although unknown (as indicated by the question mark), it is possible that LIN-65 interacts with dimethylated histones in the cytosol to be transported to the nucleus and facilitate histone exchange. Concomitantly, the histone lysine demethylases JMJD-1.2 and JMJD-3.1 promote an open chromatin state by removing methyl groups from H3K27me3, which allows the homeobox transcription factor DVE-1, with its co-activator ubiquitin-like protein 5 (UBL5), to maintain a transcriptional competent state for activating transcription factor associated with stress (ATFS-1) to bind to mitochondrial unfolded protein response elements (UPRmtE) and activate transcription of UPRmt genes. MTS, mitochondrial-targeting sequence; NLS, nuclear localization sequence.
MET-2 catalyses histone dimethylation in the cytosol before incorporation into nucleosomes in the nucleus78. It is not clear how LIN-65 nuclear accumulation is regulated, but it is possible that LIN-65 interacts with dimethylated histones in the cytosol and traffics, in complex with them, into the nucleus to facilitate histone exchange. Interestingly, nuclear localization of LIN-65 requires the mitochondrial-localized protease CLPP-1 (homologous to the bacterial protease ClpP), which is required for the activation of the UPRmt in C. elegans35, consistent with LIN-65 receiving mitochondrial-related inputs28.
In addition to a histone methyltransferase, two histone lysine demethylases containing a Jumonji C domain, JMJD-1.2 and JMJD-3.1, are also required for UPRmt activation. Presumably, the JMJD demethylases promote an open chromatin state at UPRmt genes to enable DVE-1 and ATFS-1 binding and transcription activation27, whereas LIN-65 and MET-2 mediate global chromatin compaction (Fig. 3). Chromatin remodelling during mitochondrial stress occurs independently of ATFS-1 (Ref. 28). Importantly, in the absence of ATFS-1, LIN-65 accumulates to higher levels in the nucleus, indicating that MET-2 responds to mitochondrial dysfunction via a separate, unknown mechanism28. Notably, the Jumonji methyltransferase activities require both the tricarboxylic acid (TCA) cycle product α-ketoglutarate and iron79. As the homeostasis of both molecules is affected by mitochondrial dysfunction, they may contribute to mitochondrial stress stimulation of JMJD-1.2 and JMJD-3.1.
It should be noted that overexpression of JMJD-1.2 is sufficient to activate the UPRmt, which suggests that it has an upstream role in detecting mitochondrial dysfunction and initiating the response27. However, ATFS-1 mutants lacking the MTS are also able to activate the UPRmt (Ref. 18), suggesting that the chromatin remodelling machinery and ATFS-1 are stimulated independently and then converge at the relevant UPRmt genes, where both are required. In conclusion, UPRmt activation requires at least two inputs responsive to mitochondrial activity. Chromatin remodelling stimulated by coordination of a histone dimethyltransferase and histone demethylases results in nuclear compaction while maintaining the transcriptional activation of UPRmt genes. In addition, if mitochondrial function is reduced enough to limit import of ATFS-1, the transcription factor traffics to the nucleus to activate the UPRmt. The temporal regulation of each input remains to be resolved.
UPRmt regulation via cell-to-cell communication. The UPRmt activation described above occurs in a cell-autonomous manner. However, mitochondrial dysfunction can lead to UPRmt activation in neighbouring or distant cells and tissues13,29,30,31. For example, mitochondrial perturbation in C. elegans neurons results in UPRmt activation within the neuron but also within intestinal cells, suggesting the presence of a mechanism to coordinate UPRmt activation between tissues or throughout the organism13 (Fig. 4a). Although the mechanisms of signal transduction are still under investigation, it has been proposed that it requires mitokines or other diffusible factors that are released from cells in response to mitochondrial stress to modulate UPRmt activation in other tissues13.
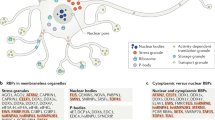
a | Mitochondrial unfolded protein response (UPRmt) regulation via cell–to–cell communication. Non-cell-autonomous UPRmt activation is initiated by mitochondrial stress in neurons, which leads to UPRmt activation in the intestine. Cell–cell communication requires serotonin secretion from serotonergic neurons and the peptide FLP-2, which is secreted from an interneuron that receives environmental inputs. The mechanism by which serotonin and FLP-2 activate the UPRmt in intestinal cells is unknown, although the receiving intestinal cell requires both activating transcription factor associated with stress (ATFS-1) and chromatin remodelling, which is mediated at least in part by the homeobox transcription factor DVE-1 and its co-activator ubiquitin-like protein 5 (UBL5). b | The role of the UPRmt in innate immunity. Bacterial pathogens that target and perturb mitochondrial function activate ATFS-1 and the UPRmt, suggesting that cells monitor mitochondrial function as a means to detect pathogenic or toxic bacteria. ATFS-1 activation in turn results in the induction of antibacterial genes to limit pathogen growth, xenobiotic detoxification genes to eliminate pathogen-derived toxins and a mitochondrial protective response to maintain and promote mitochondrial recovery.
In addition to factors that function in cell-autonomous signalling pathways, non-cell-autonomous UPRmt activation requires at least two factors that are secreted by neurons. Recently, a neuronal circuit that initiates non-cell-autonomous signalling was identified31. Serotonin, secreted from serotonergic neurons, and the peptide FLP-2, secreted from an interneuron that integrates inputs from three sensory neurons, are required for non-cell-autonomous activation (Fig. 4a). However, neither secreted molecule is sufficient to activate the UPRmt in intestinal cells29,31, indicating that additional mitokines and their receptors remain to be identified. Importantly, non-cell-autonomous signalling in the receiving intestinal cell requires both ATFS-1 and chromatin remodelling29,31 (Fig. 4a), raising the interesting question of how neuronal transmission stimulates the chromatin remodelling and transcription activation required for the UPRmt. Alternative possibilities are that the mitokine either activates each pathway independently of mitochondrial dysfunction or directly alters intestinal mitochondrial function to activate the UPRmt.
In mammals, the hormones fibroblast growth factor 21 (FGF21) and growth/differentiation factor 15 (GDF15) are induced by multiple forms of mitochondrial dysfunction and have been suggested to be mitokines30,50,51,80,81,82,83,84. FGF21 secretion is induced in an ATF4-dependent manner during mitochondrial stress and promotes β-oxidation and adipose tissue browning of white adipose tissue30,81. Induction of FGF21, along with mitochondrial chaperones and proteases, was also observed in the hearts of mice lacking DARS2 (Ref. 50). In a skeletal-muscle-specific mitochondria stress model, the induction of GDF15 depends on CHOP and regulates lipolysis and oxidative metabolism in mouse liver and adipocytes51. Importantly, the levels of FGF21 and GDF15 increase in the serum of patients with mitochondrial diseases and can serve as biomarkers of mitochondrial translation and mtDNA disorders, although FGF21 and GDF15 levels also increase in conditions not obviously related to mitochondrial stress84,85,86.
Mitochondrial recovery
Mitochondrial dysfunction induces broad changes in gene expression in all organisms examined using transcriptomic18,66 and proteomic52,87 approaches. In C. elegans, mitochondrial dysfunction caused by impairment of the mitochondrial quality control protease SPG-7, or the respiratory chain by inhibiting a complex IV component, results in altered expression of over 2,000 transcripts18,27. Less than half of these require ATFS-1 (Ref. 18); thus, the regulation of only a fraction of the cellular processes affected by UPRmt activation is understood, which suggests that additional transcription factors are activated during mitochondrial dysfunction88. Below, we examine the transcriptional outputs of the UPRmt mediated by known transcription factors and the interactions with other mitochondrial stress response pathways.
UPRmt transcriptional outputs. The conditions that activate the UPRmt also interfere with mitochondrial protein import. For example, OXPHOS perturbation, excessive ROS and mitonuclear protein imbalance (owing to incorrect protein folding or complex assembly) in the matrix or inner membrane all interfere with protein import (Fig. 1). In response, ATFS-1 activates many genes, some of which function to repair these defects (Table 2). ATFS-1 binds to a conserved promoter element (UPRmtE; very similar to an amino acid response element) in many of these genes required for their induction89 (Fig. 3). Of note, ATF4 and ATF5 bind to the same promoter element64,66,89. To re-establish mitochondrial proteostasis, mitochondrial chaperone and quality control protease genes are induced to facilitate de novo protein folding and degrade those proteins that have become damaged or fail to fold. More specifically related to OXPHOS activity, OXPHOS complex assembly factors, components required for iron–sulfur cluster biogenesis and most mitochondrial ribosome subunits are also induced. In addition, multiple components that limit ROS toxicity, including a superoxide dismutase, glutathione synthesis machinery and ubiquinone synthesis genes, are also induced. Finally, the UPRmt promotes import recovery by inducing transcription of the core components of the TIM23 complex required to import mitochondrial-targeted proteins into the matrix18.
UPRmt activation also results in metabolic remodelling89. ATFS-1 temporarily limits OXPHOS gene transcription in both the nucleus and the mitochondria while simultaneously increasing transcription of all glycolysis components18,89. This shift promotes mitochondrial recovery by temporarily reducing expression of the most highly expressed and difficult to process mitochondrial proteins to limit the protein folding burden on the stressed compartment. Increased glycolysis probably allows cells to generate ATP in the cytosol to promote survival and mitochondrial recovery89,18. Additionally, loss of the TCA cycle enzyme fumarate hydratase, mitochondrial (FH1; also known as FH), which catalyses the hydration of fumarate to malate, induces ATF4 (Ref. 66) and the UPRmt in mammals and C. elegans90, consistent with metabolic perturbations activating the UPRmt, which in turn adapts metabolism. Furthermore, mitochondrial dysfunction in flies and mammalian cells was recently found to remodel one-carbon metabolism, which promotes cellular redox balance and nucleotide synthesis66,72,91. In human cells, mitochondrial dysfunction impaired the serine-dependent synthesis of formate and led to an ATF4-dependent increase in the biosynthesis of serine and transsulfuration in human cells72. Importantly, formate or purine supplementation was able to rescue growth and the defects caused by OXPHOS inhibitors, emphasizing the importance of these findings72.
Once mitochondrial function and protein import are restored, UPRmt activity is reduced as ATFS-1 is again efficiently imported into mitochondria and degraded. Thus, because of the short half-life of ATFS-1, the UPRmt can respond to transient mitochondrial perturbations.
Pathways complementary to the UPRmt contribute to organelle recovery. To restore mitochondrial proteostasis and organelle recovery, transcriptional adaptations occur at the same time as a reduction in protein synthesis. In both mammals and C. elegans, mitochondrial stress leads to activation of the kinase GCN2, which phosphorylates the translation initiation factor eIF2α to reduce global protein synthesis45,92 (Fig. 2). This function reduces the influx of newly synthesized proteins into mitochondria so as not to overwhelm the proteostasis machinery and facilitates protein folding and complex assembly45. eIF2α phosphorylation in combination with increased transcription mediated by ATFS-1, ATF4 and ATF5 may also prioritize the import of proteins to re-establish proteostasis and adapt metabolism.
A recent study demonstrated that a specific mitochondrial perturbation in C. elegans, caused by inhibition of the mitochondrial chaperone HSP-6 or fatty acid oxidation, led to simultaneous activation of both the UPRmt and the heat shock response (HSR)93. The activation of the HSR under these conditions was dependent on the activation of the UPRmt and could reduce proteotoxicity caused by poly-glutamine aggregates93. These findings suggest that the UPRmt can interact with other stress responses to coordinate the appropriate response to resolve a specific condition93.
The UPRmt and mitochondrial-specific autophagy (mitophagy), which eliminates severely damaged mitochondria, can be activated simultaneously, as both are responsive to similar forms of mitochondrial dysfunction25,26,94, including damage by paraquat38,95, mtDNA mutations or deletions25,26,96,97 and unfolded protein accumulation in the matrix20,98. The mitophagy component PTEN-induced putative kinase 1 (PINK1) is imported into healthy mitochondria, processed and targeted for degradation99. However, during mitochondrial stress, PINK1 fails to be imported across the inner membrane and instead accumulates on the mitochondrial outer membrane of defective mitochondria95,98,100,101. Here, PINK1 phosphorylates ubiquitin and the ubiquitin ligase parkin, leading to the engulfment of the defective organelle by autophagosomes and subsequent degradation within lysosomes95,99,102,103. The events orchestrated by the UPRmt and mitophagy are fairly clear (transcription adaptation to promote recovery of mitochondrial activity versus degradation of severely damaged mitochondria); however, how they are coordinated to recover the mitochondrial network remains to be determined.
Implications in physiology and ageing
UPRmt activation is essential to ensure normal development in the event of mitochondrial stress or OXPHOS dysfunction, consistent with a role of the UPRmt in inducing metabolic adaptations and a mitochondrial recovery programme18,45,89. UPRmt activation is strong during development but relatively mild during adulthood13,32. However, paradoxically, mild mitochondrial perturbations that activate the UPRmt during development result in increased longevity in worms, flies and mice32,104,105,106,107,108,109,110,111,112. Conversely, mitochondrial perturbations during adulthood do not extend lifespan or activate the UPRmt, suggesting that the pathway is differentially regulated as organisms age13,32,33. It is not clear whether adult cells have reduced capability to detect mitochondrial stress, but it was found that chromatin compaction and remodelling occur near the onset of adulthood and that these chromatin changes limit UPRmt activation during ageing27,28,113. Below, we examine physiological roles for UPRmt signalling during conditions affected by ageing.
The UPRmt and increased longevity. As stated above, there is a strong correlation between mitochondrial defects that extend lifespan and those that activate the UPRmt. For example, modest perturbations of OXPHOS complex IV or the mitochondrial ribosome extend lifespan and activate the UPRmt in worms and mice13,15,32,114. However, perturbation of OXPHOS complex II (succinate dehydrogenase) activates the UPRmt but does not extend lifespan in C. elegans13,114,115,116. Multiple studies indicate that the UPRmt has a role in extending lifespan. For example, inhibition of ATFS-1, the DVE-1 co-activator ubiquitin-like protein 5 (UBL5) and the Jumonji histone demethylases reduces longevity during mitochondrial stress13,15,27,117. Furthermore, overexpression of the histone demethylases is sufficient to activate the UPRmt and extend lifespan27. Mitochondrial dysfunction or overexpression of JMJD-1.2 is sufficient to delay chromatin compaction, allowing UPRmt activation into adulthood27.
Additional mitochondrial stress response pathways are required for increased longevity, consistent with mitochondrial dysfunction being pleiotropic. Hypoxia-inducible factor 1 (HIF-1) is required for lifespan extension during mitochondrial stress, as are several MAP kinases in C. elegans88,118,119. Mitochondrial-generated ROS also increase longevity in a manner dependent on the programmed cell death machinery in C. elegans112. Although several signalling components that promote longevity following mitochondrial dysfunction have been identified, the cellular processes and physiological changes that are involved remain to be defined. Is metabolic remodelling and/or the activation of genes that limit damage during ageing the cause of increased longevity120? Although the longevity-promoting cellular processes affected by the UPRmt are unclear, it is known that they require chromatin remodelling induced by the demethylases JMJD-1.2 and/or JMJD-1.3, which maintain a chromatin state necessary for UPRmt activation27.
The UPRmt in physiology and disease. The UPRmt seems to be associated with diverse pathologies, which is consistent with mitochondrial dysfunction being common to many diseases. For example, increased levels of transcripts associated with UPRmt activation have been observed in patients with myopathy caused by mitochondrial disorders85, in patients with either sporadic or familial Alzheimer disease121, in a mouse model of cardiomyopathy caused by loss of a mitochondrial tRNA synthetase50, in a mouse model of depression caused by chronic restraint122 and in glioblastomas123. Although these observations are consistent with the hypothesis that the UPRmt becomes activated, the transcription factors required for the observed mitochondrial-stress-induced transcription remain to be determined in each pathological scenario, as do the effects of the transcriptional response on physiology.
Recent evidence suggests that UPRmt transcripts can function as biomarkers of mitochondrial dysfunction or diseases caused by mitochondrial dysfunction and provide potential therapeutic approaches to treat such conditions. For example, the hormones FGF21 (Ref. 85) and GDF15 (Ref. 51) are biomarkers of myopathy caused by mitochondrial dysfunction. Treatments that increase NAD+ activate the UPRmt and ameliorate mitochondrial function in aged tissues to promote muscle stem cell maintenance and regeneration124,125. Moreover, administration of GDF15 improved insulin sensitivity in obese mice12, and supplementation with an NAD+ precursor alleviated non-alcoholic fatty liver disease126, although it remains to be determined if these improvements are due to UPRmt activation.
Although the UPRmt is a stress response, low levels of constitutive UPRmt activation have been observed and potentially contribute to mitochondrial maintenance during normal development. For example, a recent comparative study of 40 genetically diverse mouse strains revealed a coordinated expression pattern of conserved UPRmt components, including mitochondrial chaperones and proteases, that negatively correlated with expression of the mitochondrial quality control protease SPG7 in both mice and humans87. These data are consistent with SPG-7 inhibition strongly activating the UPRmt in C. elegans18,19. These studies suggest that the UPRmt buffers against more subtle forms of mitochondrial stress that occur in normal populations simply due to genotypic variance. Additionally, mice lacking ATF4 (Ref. 127) or ATF5 (Refs 128,129) have numerous developmental defects consistent with the UPRmt having baseline functions during normal development.
The UPRmt as an innate immune response. Both adaptive and innate immunity decline with age and rely on mitochondria for proper regulation130,131. Mitochondria are signalling platforms for innate immune pathways that are responsive to bacterial and viral infections and produce ROS to eliminate intracellular bacteria130. Mitochondria are not only immune response hubs but also targets of pathogen virulence factors132. Interestingly, mitochondrial perturbation results in cellular stress responses tightly associated with innate immunity (Fig. 4b). For example, herpesvirus infection results in the release of mtDNA into the cytosol, which activates an antiviral response130,133. In addition, the bacterium Helicobacter pylori produces a pore-forming protein targeted to mitochondria that perturbs OXPHOS134. Similarly, Pseudomonas aeruginosa secretes the OXPHOS inhibitor cyanide and iron-chelating proteins as virulence factors135. These observations led to the intriguing hypothesis that monitoring mitochondrial function enables cells to detect pathogens, as mitochondria seem to be common targets of virulence factors136.
Recent studies reported that exposure to multiple bacterial species or bacterial-derived toxins activates the UPRmt (Refs 16,24). Importantly, in addition to a mitochondrial protective transcriptional programme, ATFS-1 induces transcription of multiple antibacterial genes, including antimicrobial peptides and secreted lysozymes, as well as a xenobiotic detoxification programme, suggesting a role in pathogen detection and resistance to toxin exposure16,24 (Fig. 4b). Exposure to P. aeruginosa activates the UPRmt, which requires P. aeruginosa to produce cyanide and iron chelators, suggesting that monitoring of mitochondrial function is used to activate an innate immune response. Furthermore, animals lacking ATFS-1 are susceptible to P. aeruginosa accumulation in the intestine, resulting in earlier death. However, hyperactivation of the UPRmt provides resistance to the pathogen, both slowing intestinal colonization and extending animal survival time, indicating that the UPRmt is a bona fide innate immune response16,24,137.
Although the virulence factors that perturb mitochondrial function are largely unknown, a recent survey of natural bacterial isolates revealed that many species cause UPRmt activation, suggesting that mechanisms to perturb mitochondria are found in many bacterial species and may be widespread16. It should be noted that the OXPHOS inhibitors antimycin and oligomycin are derived from the bacteria Streptomyces, as is the mitochondrial ribosome inhibitor tetracycline138,139. While Streptomyces is not a human pathogen, it likely uses these toxins to impair competing bacterial species in the soil where it lives. This suggests that the OXPHOS toxins that evolved to impair competitive bacterial species are also useful virulence factors against mitochondria, which themselves are bacterial-derived.
The UPRmt and the propagation of deleterious mitochondrial genomes. As discussed above, the decline in UPRmt activation with ageing potentially contributes to age-associated cell and tissue dysfunction. Thus, approaches to prolong UPRmt activation during ageing may alleviate age-associated cell dysfunction. However, recent work has indicated that prolonged UPRmt activation has negative consequences. In addition to reduced fecundity39, prolonged UPRmt activation promotes, or facilitates, the propagation of deleterious mtDNAs in germ cells and somatic cells, suggesting that the decline in UPRmt activation during ageing may have been evolutionarily selected25,26 (Fig. 5a).
a | The presence of both wild-type (WT) mitochondrial genomes (mtDNA; blue) and deleterious mutant mtDNAs (ΔmtDNA; red) in the same cell is referred to as heteroplasmy. Accumulation of ΔmtDNA can cause mitochondrial diseases as well as age-associated damage to cells such as muscles and neurons. ΔmtDNA accumulation leads to oxidative phosphorylation dysfunction, which activates activating transcription factor associated with stress (ATFS-1) and the mitochondrial unfolded protein response (UPRmt). By inducing a mitochondrial recovery programme that includes mitochondrial biogenesis as well as mitochondrial fission and fusion genes, the UPRmt promotes the preferential propagation of the ΔmtDNA at the expense of the wild-type mtDNA. Severely damaged mitochondria that contain high levels of ΔmtDNA are degraded by mitophagy. b | Inhibition of ATFS-1 and the UPRmt results in preferential loss of the ΔmtDNA while maintaining the wild-type mtDNA.
Most somatic cells contain hundreds of mtDNAs that accumulate mutations at much higher rates than nuclear genes140. Mutations or deletions in individual mtDNAs are well tolerated because of the high copy number of mtDNAs in each cell. However, if the percentage of mtDNAs carrying a specific deleterious mutation reaches ∼60–80%, depending on the severity of the loss-of-function mutation and the tissue type, mtDNA mutations can lead to mitochondrial dysfunction, compromising cellular function141. One mechanism to acquire a high load of ΔmtDNAs (mtDNAs carrying deletions) is through inheritance via the maternal germ line. However, deep sequencing data from otherwise healthy individuals have indicated that ΔmtDNAs are commonly found in multiple tissues141,142. But as the number of mutated mtDNAs is usually quite low, cellular function remains unaffected. However, during ageing, a ΔmtDNA clone can outcompete wild-type mtDNAs, resulting in high levels of heteroplasmy, typically in muscle cells or neurons. The accumulation of ΔmtDNAs during ageing and their damaging effects on cellular function are well-documented and have been reviewed recently143. Below, we focus on the interactions between ΔmtDNAs and UPRmt activation.
The mechanisms that promote the preferential accumulation of ΔmtDNAs in somatic cells are not well understood owing at least in part to the availability of only a few metazoan models that stably inherit easily tractable ΔmtDNAs. However, a heteroplasmic C. elegans strain harbouring ∼60% ΔmtDNAs with a 3-kilobase deletion has provided insight into the maintenance of deleterious heteroplasmies144. Perhaps not surprisingly, the absence of four essential OXPHOS genes from 60% of the mtDNAs perturbs OXPHOS function and causes UPRmt activation. UPRmt activation caused by ΔmtDNAs protects against OXPHOS perturbations in a similar manner to the UPRmt, protecting cells from mutations in nuclear-encoded OXPHOS genes or toxins that impair OXPHOS18. Surprisingly, ATFS-1 inhibition caused a near complete loss of the deleterious mtDNA, indicating that UPRmt activation can preferentially propagate or maintain the deleterious mtDNA (Fig. 5b). Alternatively, further UPRmt activation resulted in increased accumulation of the ΔmtDNA25,26. Accumulation of ΔmtDNAs via UPRmt activation is antagonized by mitophagy, as mitophagy impairment allows further ΔmtDNA accumulation25,26. Presumably, as the heteroplasmy ratio increases, so does mitochondrial damage, resulting in selective degradation of those mitochondria with the highest ratio of ΔmtDNAs.
One possible mechanism by which the UPRmt promotes or allows ΔmtDNA accumulation is by mitigating mitochondrial damage, allowing ΔmtDNAs to 'escape' mitophagy. Alternatively, the UPRmt-mediated mitochondrial recovery programme may more directly promote the propagation of the ΔmtDNA25,26. Because the UPRmt responds to a diverse variety of mitochondrial stressors, the repair and recovery programme is not necessarily specific to the underlying cause of mitochondrial stress. For example, exposure to OXPHOS inhibitors elicits a UPRmt that includes mtDNA replication as part of the recovery programme18. Similarly, OXPHOS perturbation by ΔmtDNAs also elicits a UPRmt, but in this case the ΔmtDNA may be preferentially replicated during organelle recovery.
Conclusions and perspectives
New and unexpected roles have been uncovered for the UPRmt that extend far beyond its original identified role in restoring mitochondrial proteostasis. The rapid rate at which these discoveries have been made suggests that the field will continue to evolve to include additional modes of regulation and physiological roles. Multiple UPRmt signalling components involved in direct transcriptional adaptations and chromatin remodelling have been found that affect age-associated mitochondrial function. While activation of the UPRmt is potentially beneficial and may serve as a future therapeutic target, the impact of activating this response long term is not fully understood and should be undertaken cautiously, as prolonged UPRmt activation can lead to the propagation of mitochondrial damage25,26,29. Additionally, many of the genes induced by the UPRmt are of unknown function and should be characterized along with additional transcription factors required to regulate the full transcriptional response to mitochondrial dysfunction18,88.
The regulation of the UPRmt in mammals is potentially more complex than that described in C. elegans. In addition, while a mammalian orthologue of ATFS-1 was recently identified64, the requirement for at least two additional transcription factors, ATF4 and CHOP, along with four eIF2α kinases suggests that much remains to be determined. For example, which eIF2α kinases respond to mitochondrial dysfunction and how do they do so? The discovery that UPRmt activation can be transmitted between cells capable of coordinating tissues and protective effects throughout the organism has suggested the existence of mitokines. While the mechanism by which the UPRmt is communicated is only beginning to be understood, the identification of additional extracellular components will undoubtedly have therapeutic implications as well as contribute to the understanding of the molecular mechanisms that regulate and coordinate the UPRmt.
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Gray, M. W., Burger, G. & Lang, B. F. The origin and early evolution of mitochondria. Genome Biol. 2, 1018.1–1018.5 (2001).
Lane, N. & Martin, W. The energetics of genome complexity. Nature 467, 929–934 (2010).
Mishra, P. & Chan, D. C. Mitochondrial dynamics and inheritance during cell division, development and disease. Nat. Rev. Mol. Cell Biol. 15, 634–646 (2014).
Schmidt, O., Pfanner, N. & Meisinger, C. Mitochondrial protein import: from proteomics to functional mechanisms. Nat. Rev. Mol. Cell Biol. 11, 655–667 (2010).
Nunnari, J. & Suomalainen, A. Mitochondria: in sickness and in health. Cell 148, 1145–1159 (2012).
Suomalainen A. & Battersby, B. J. Mitochondrial diseases: the contribution of organelle stress responses to pathology. Nat. Rev. Mol. Cell Biol. http://dx.doi.org/10.1038/nrm.2017.66 (2017).
Sun, N., Youle, R. J. & Finkel, T. The mitochondrial basis of aging. Mol. Cell 61, 654–666 (2016).
Bratic, A. & Larsson, N. G. The role of mitochondria in aging. J. Clin. Invest. 123, 951–957 (2013).
Morris, A. A. et al. Deficiency of respiratory chain complex I is a common cause of Leigh disease. Ann. Neurol. 40, 25–30 (1996).
Wallace, D. C. et al. Mitochondrial DNA mutation associated with Leber's hereditary optic neuropathy. Science 242, 1427–1430 (1988).
Peterson, C. M., Johannsen, D. L. & Ravussin, E. Skeletal muscle mitochondria and aging: a review. J. Aging Res. 2012, 194821 (2012).
Schulz, A. M. & Haynes, C. M. UPRmt-mediated cytoprotection and organismal aging. Biochim. Biophys. Acta 1847, 1448–1456 (2015).
Durieux, J., Wolff, S. & Dillin, A. The cell-non-autonomous nature of electron transport chain-mediated longevity. Cell 144, 79–91 (2011). This study reveals that mitochondrial UPRmt activation in neurons can be communicated to cells in different tissues to mediate longevity.
Haynes, C. M., Yang, Y., Blais, S. P., Neubert, T. A. & Ron, D. The matrix peptide exporter HAF-1 signals a mitochondrial UPR by activating the transcription factor ZC376.7 in C. elegans. Mol. Cell 37, 529–540 (2010).
Houtkooper, R. H. et al. Mitonuclear protein imbalance as a conserved longevity mechanism. Nature 497, 451–457 (2013). This study indicates that an imbalance between nuclear-encoded and mitochondrial-encoded OXPHOS proteins induces the UPRmt and extends lifespan in mice and worms.
Liu, Y., Samuel, B. S., Breen, P. C. & Ruvkun, G. Caenorhabditis elegans pathways that surveil and defend mitochondria. Nature 508, 406–410 (2014). This study reports that mitochondrial dysfunction causes induction of detoxification and innate immune genes.
Martinus, R. D. et al. Selective induction of mitochondrial chaperones in response to loss of the mitochondrial genome. Eur. J. Biochem. 240, 98–103 (1996).
Nargund, A. M., Pellegrino, M. W., Fiorese, C. J., Baker, B. M. & Haynes, C. M. Mitochondrial import efficiency of ATFS-1 regulates mitochondrial UPR activation. Science 337, 587–590 (2012). This study suggests how mitochondrial dysfunction is sensed and communicated to the nucleus to induce a broad transcriptional response.
Yoneda, T. et al. Compartment-specific perturbation of protein handling activates genes encoding mitochondrial chaperones. J. Cell Sci. 117, 4055–4066 (2004).
Zhao, Q. et al. A mitochondrial specific stress response in mammalian cells. EMBO J. 21, 4411–4419 (2002).
Chan, D. C. Fusion and fission: interlinked processes critical for mitochondrial health. Annu. Rev. Genet. 46, 265–287 (2012).
Lewis, S. C., Uchiyama, L. F. & Nunnari, J. ER-mitochondria contacts couple mtDNA synthesis with mitochondrial division in human cells. Science 353, aaf5549 (2016).
Lin, Y. F. & Haynes, C. M. Metabolism and the UPRmt. Mol. Cell 61, 677–682 (2016).
Pellegrino, M. W. et al. Mitochondrial UPR-regulated innate immunity provides resistance to pathogen infection. Nature 516, 414–417 (2014). This study shows that the UPRmt can serve as a means to detect pathogens that perturb mitochondria and provide an antibacterial response.
Gitschlag, B. L. et al. Homeostatic responses regulate selfish mitochondrial genome dynamics in C. elegans. Cell Metab. 24, 91–103 (2016).
Lin, Y. F. et al. Maintenance and propagation of a deleterious mitochondrial genome by the mitochondrial unfolded protein response. Nature 533, 416–419 (2016). References 25 and 26 report that UPRmt activation preferentially maintains a ΔmtDNA in C. elegans.
Merkwirth, C. et al. Two conserved histone demethylases regulate mitochondrial stress-induced longevity. Cell 165, 1209–1223 (2016). This study reveals a role for two histone demethylases and chromatin remodelling in the UPRmt.
Tian, Y. et al. Mitochondrial stress induces chromatin reorganization to promote longevity and UPRmt. Cell 165, 1197–1208 (2016). This study shows that mitochondrial stress results in nuclear compaction with concomitant opening up of UPRmt chromatin regions.
Berendzen, K. M. et al. Neuroendocrine coordination of mitochondrial stress signaling and proteostasis. Cell 166, 1553–1563.e10 (2016).
Kim, K. H. et al. Autophagy deficiency leads to protection from obesity and insulin resistance by inducing Fgf21 as a mitokine. Nat. Med. 19, 83–92 (2013).
Shao, L. W., Niu, R. & Liu, Y. Neuropeptide signals cell non-autonomous mitochondrial unfolded protein response. Cell Res. 26, 1182–1196 (2016).
Dillin, A. et al. Rates of behavior and aging specified by mitochondrial function during development. Science 298, 2398–2401 (2002).
Rea, S. L., Ventura, N. & Johnson, T. E. Relationship between mitochondrial electron transport chain dysfunction, development, and life extension in Caenorhabditis elegans. PLoS Biol. 5, e259 (2007).
Walter, P. & Ron, D. The unfolded protein response: from stress pathway to homeostatic regulation. Science 334, 1081–1086 (2011).
Haynes, C. M., Petrova, K., Benedetti, C., Yang, Y. & Ron, D. ClpP mediates activation of a mitochondrial unfolded protein response in C. elegans. Dev. Cell 13, 467–480 (2007).
Bennett, C. F. et al. Activation of the mitochondrial unfolded protein response does not predict longevity in Caenorhabditis elegans. Nat. Commun. 5, 3483 (2014).
Benedetti, C., Haynes, C. M., Yang, Y., Harding, H. P. & Ron, D. Ubiquitin-like protein 5 positively regulates chaperone gene expression in the mitochondrial unfolded protein response. Genetics 174, 229–239 (2006).
Runkel, E. D., Liu, S., Baumeister, R. & Schulze, E. Surveillance-activated defenses block the ROS-induced mitochondrial unfolded protein response. PLoS Genet. 9, e1003346 (2013).
Rauthan, M., Ranji, P., Aguilera Pradenas, N., Pitot, C. & Pilon, M. The mitochondrial unfolded protein response activator ATFS-1 protects cells from inhibition of the mevalonate pathway. Proc. Natl Acad. Sci. USA 110, 5981–5986 (2013).
Williams, C. C., Jan, C. H. & Weissman, J. S. Targeting and plasticity of mitochondrial proteins revealed by proximity-specific ribosome profiling. Science 346, 748–751 (2014).
Harbauer, A. B., Zahedi, R. P., Sickmann, A., Pfanner, N. & Meisinger, C. The protein import machinery of mitochondria-a regulatory hub in metabolism, stress, and disease. Cell Metab. 19, 357–372 (2014).
Wright, G., Terada, K., Yano, M., Sergeev, I. & Mori, M. Oxidative stress inhibits the mitochondrial import of preproteins and leads to their degradation. Exp. Cell Res. 263, 107–117 (2001).
Kang, P. J. et al. Requirement for hsp70 in the mitochondrial matrix for translocation and folding of precursor proteins. Nature 348, 137–143 (1990).
Neupert, W. & Brunner, M. The protein import motor of mitochondria. Nat. Rev. Mol. Cell Biol. 3, 555–565 (2002).
Baker, B. M., Nargund, A. M., Sun, T. & Haynes, C. M. Protective coupling of mitochondrial function and protein synthesis via the eIF2α kinase GCN-2. PLoS Genet. 8, e1002760 (2012).
Stiburek, L. et al. YME1L controls the accumulation of respiratory chain subunits and is required for apoptotic resistance, cristae morphogenesis, and cell proliferation. Mol. Biol. Cell 23, 1010–1023 (2012).
Martinelli, P. & Rugarli, E. I. Emerging roles of mitochondrial proteases in neurodegeneration. Biochim. Biophys. Acta 1797, 1–10 (2010).
Wrobel, L. et al. Mistargeted mitochondrial proteins activate a proteostatic response in the cytosol. Nature 524, 485–488 (2015).
Wang, X. & Chen, X. J. A cytosolic network suppressing mitochondria-mediated proteostatic stress and cell death. Nature 524, 481–484 (2015).
Dogan, S. A. et al. Tissue-specific loss of DARS2 activates stress responses independently of respiratory chain deficiency in the heart. Cell Metab. 19, 458–469 (2014). This is one of the first in vivo mammalian studies demonstrating that mitochondrial dysfunction in the heart leads to the activation of the ISR.
Chung, H. K. et al. Growth differentiation factor 15 is a myomitokine governing systemic energy homeostasis. J. Cell Biol. 216, 149–165 (2017).
Munch, C. & Harper, J. W. Mitochondrial unfolded protein response controls matrix pre-RNA processing and translation. Nature 534, 710–713 (2016).
Harding, H. P. et al. An integrated stress response regulates amino acid metabolism and resistance to oxidative stress. Mol. Cell 11, 619–633 (2003).
Teske, B. F. et al. CHOP induces activating transcription factor 5 (ATF5) to trigger apoptosis in response to perturbations in protein homeostasis. Mol. Biol. Cell 24, 2477–2490 (2013).
Pakos-Zebrucka, K. et al. The integrated stress response. EMBO Rep. 17, 1374–1395 (2016).
Dey, S. et al. Both transcriptional regulation and translational control of ATF4 are central to the integrated stress response. J. Biol. Chem. 285, 33165–33174 (2010).
Zhang, P. et al. The GCN2 eIF2α kinase is required for adaptation to amino acid deprivation in mice. Mol. Cell. Biol. 22, 6681–6688 (2002).
Yan, W. et al. Control of PERK eIF2α kinase activity by the endoplasmic reticulum stress-induced molecular chaperone P58IPK. Proc. Natl Acad. Sci. USA 99, 15920–15925 (2002).
Hori, O. et al. Transmission of cell stress from endoplasmic reticulum to mitochondria: enhanced expression of Lon protease. J. Cell Biol. 157, 1151–1160 (2002).
Harding, H. P. et al. Regulated translation initiation controls stress-induced gene expression in mammalian cells. Mol. Cell 6, 1099–1108 (2000).
Zhou, D. et al. Phosphorylation of eIF2 directs ATF5 translational control in response to diverse stress conditions. J. Biol. Chem. 283, 7064–7073 (2008).
Jousse, C. et al. Inhibition of CHOP translation by a peptide encoded by an open reading frame localized in the chop 5'UTR. Nucleic Acids Res. 29, 4341–4351 (2001).
Aldridge, J. E., Horibe, T. & Hoogenraad, N. J. Discovery of genes activated by the mitochondrial unfolded protein response (mtUPR) and cognate promoter elements. PLoS ONE 2, e874 (2007).
Fiorese, C. J. et al. The transcription factor ATF5 mediates a mammalian mitochondrial UPR. Curr. Biol. 26, 2037–2043 (2016).
Horibe, T. & Hoogenraad, N. J. The chop gene contains an element for the positive regulation of the mitochondrial unfolded protein response. PLoS ONE 2, e835 (2007).
Quiros, P. M. et al. Multi-omics analysis identifies ATF4 as a key regulator of the mitochondrial stress response in mammals. J. Cell Biol. 216, 2027–2045 (2017).
Fawcett, T. W., Martindale, J. L., Guyton, K. Z., Hai, T. & Holbrook, N. J. Complexes containing activating transcription factor (ATF)/cAMP-responsive-element-binding protein (CREB) interact with the CCAAT/enhancer-binding protein (C/EBP)-ATF composite site to regulate Gadd153 expression during the stress response. Biochem. J. 339, 135–141 (1999).
Wang, X. Z. et al. Signals from the stressed endoplasmic reticulum induce C/EBP-homologous protein (CHOP/GADD153). Mol. Cell. Biol. 16, 4273–4280 (1996).
Fraser, J. E., Wang, C., Chan, K. W., Vasudevan, S. G. & Jans, D. A. Novel dengue virus inhibitor 4-HPR activates ATF4 independent of Protein Kinase R-like Endoplasmic Reticulum Kinase and elevates levels of eIF2α phosphorylation in virus infected cells. Antiviral Res. 130, 1–6 (2016).
Watatani, Y. et al. Stress-induced translation of ATF5 mRNA is regulated by the 5′-untranslated region. J. Biol. Chem. 283, 2543–2553 (2008).
Lee, Y. Y., Cevallos, R. C. & Jan, E. An upstream open reading frame regulates translation of GADD34 during cellular stresses that induce eIF2α phosphorylation. J. Biol. Chem. 284, 6661–6673 (2009).
Bao, X. R. et al. Mitochondrial dysfunction remodels one-carbon metabolism in human cells. eLife 5, e10575 (2016).
Fusakio, M. E. et al. Transcription factor ATF4 directs basal and stress-induced gene expression in the unfolded protein response and cholesterol metabolism in the liver. Mol. Biol. Cell 27, 1536–1551 (2016).
Marciniak, S. J. et al. CHOP induces death by promoting protein synthesis and oxidation in the stressed endoplasmic reticulum. Genes Dev. 18, 3066–3077 (2004).
Radke, S. et al. Mitochondrial protein quality control by the proteasome involves ubiquitination and the protease Omi. J. Biol. Chem. 283, 12681–12685 (2008).
Papa, L. & Germain, D. Estrogen receptor mediates a distinct mitochondrial unfolded protein response. J. Cell Sci. 124, 1396–1402 (2011).
Riar, A. K. et al. Sex specific activation of the ERα axis of the mitochondrial UPR (UPRmt) in the G93A-SOD1 mouse model of familial ALS. Hum. Mol. Genet. 26, 1318–1327 (2017).
Towbin, B. D. et al. Step-wise methylation of histone H3K9 positions heterochromatin at the nuclear periphery. Cell 150, 934–947 (2012).
Hoffmann, I. et al. The role of histone demethylases in cancer therapy. Mol. Oncol. 6, 683–703 (2012).
Kim, K. H. et al. Metformin-induced inhibition of the mitochondrial respiratory chain increases FGF21 expression via ATF4 activation. Biochem. Biophys. Res. Commun. 440, 76–81 (2013).
Keipert, S. et al. Skeletal muscle mitochondrial uncoupling drives endocrine cross-talk through the induction of FGF21 as a myokine. Am. J. Physiol. Endocrinol. Metab. 306, E469–E482 (2014).
Kim, K. H. & Lee, M. S. FGF21 as a stress hormone: the roles of FGF21 in stress adaptation and the treatment of metabolic diseases. Diabetes Metab. J. 38, 245–251 (2014).
Gleyzer, N. & Scarpulla, R. C. PGC-1-related coactivator (PRC), a sensor of metabolic stress, orchestrates a redox-sensitive program of inflammatory gene expression. J. Biol. Chem. 286, 39715–39725 (2011).
Lehtonen, J. M. et al. FGF21 is a biomarker for mitochondrial translation and mtDNA maintenance disorders. Neurology 87, 2290–2299 (2016).
Suomalainen, A. et al. FGF-21 as a biomarker for muscle-manifesting mitochondrial respiratory chain deficiencies: a diagnostic study. Lancet Neurol. 10, 806–818 (2011).
Fujita, Y. et al. GDF15 is a novel biomarker to evaluate efficacy of pyruvate therapy for mitochondrial diseases. Mitochondrion 20, 34–42 (2015).
Wu, Y. et al. Multilayered genetic and omics dissection of mitochondrial activity in a mouse reference population. Cell 158, 1415–1430 (2014).
Munkacsy, E. et al. DLK-1, SEK-3 and PMK-3 are required for the life extension induced by mitochondrial bioenergetic disruption in C. elegans. PLoS Genet. 12, e1006133 (2016).
Nargund, A. M., Fiorese, C. J., Pellegrino, M. W., Deng, P. & Haynes, C. M. Mitochondrial and nuclear accumulation of the transcription factor ATFS-1 promotes OXPHOS recovery during the UPRmt. Mol. Cell 58, 123–133 (2015).
Wang, X. et al. Joint mouse-human phenome-wide association to test gene function and disease risk. Nat. Commun. 7, 10464 (2016).
Celardo, I., Lehmann, S., Costa, A. C., Loh, S. H. & Miguel Martins, L. dATF4 regulation of mitochondrial folate-mediated one-carbon metabolism is neuroprotective. Cell Death Differ. 24, 638–648 (2017).
Michel, S., Canonne, M., Arnould, T. & Renard, P. Inhibition of mitochondrial genome expression triggers the activation of CHOP-10 by a cell signaling dependent on the integrated stress response but not the mitochondrial unfolded protein response. Mitochondrion 21, 58–68 (2015).
Kim, H. E. et al. Lipid biosynthesis coordinates a mitochondrial-to-cytosolic stress response. Cell 166, 1539–1552.e16 (2016).
Georgakopoulos, N. D., Wells, G. & Campanella, M. The pharmacological regulation of cellular mitophagy. Nat. Chem. Biol. 13, 136–146 (2017).
Narendra, D. P. et al. PINK1 is selectively stabilized on impaired mitochondria to activate Parkin. PLoS Biol. 8, e1000298 (2010).
Pickrell, A. M. et al. Endogenous Parkin preserves dopaminergic substantia nigral neurons following mitochondrial DNA mutagenic stress. Neuron 87, 371–381 (2015).
Suen, D. F., Narendra, D. P., Tanaka, A., Manfredi, G. & Youle, R. J. Parkin overexpression selects against a deleterious mtDNA mutation in heteroplasmic cybrid cells. Proc. Natl Acad. Sci. USA 107, 11835–11840 (2010).
Jin, S. M. & Youle, R. J. The accumulation of misfolded proteins in the mitochondrial matrix is sensed by PINK1 to induce PARK2/Parkin-mediated mitophagy of polarized mitochondria. Autophagy 9, 1750–1757 (2013).
Youle, R. J. & Narendra, D. P. Mechanisms of mitophagy. Nat. Rev. Mol. Cell Biol. 12, 9–14 (2011).
Burbulla, L. F. et al. Mitochondrial proteolytic stress induced by loss of mortalin function is rescued by Parkin and PINK1. Cell Death Dis. 5, e1180 (2014).
Geisler, S. et al. PINK1/Parkin-mediated mitophagy is dependent on VDAC1 and p62/SQSTM1. Nat. Cell Biol. 12, 119–131 (2010).
Vives-Bauza, C. et al. PINK1-dependent recruitment of Parkin to mitochondria in mitophagy. Proc. Natl Acad. Sci. USA 107, 378–383 (2010).
Kim, Y. et al. PINK1 controls mitochondrial localization of Parkin through direct phosphorylation. Biochem. Biophys. Res. Commun. 377, 975–980 (2008).
Felkai, S. et al. CLK-1 controls respiration, behavior and aging in the nematode Caenorhabditis elegans. EMBO J. 18, 1783–1792 (1999).
Feng, J., Bussiere, F. & Hekimi, S. Mitochondrial electron transport is a key determinant of life span in Caenorhabditis elegans. Dev. Cell 1, 633–644 (2001).
Lee, S. S. et al. A systematic RNAi screen identifies a critical role for mitochondria in C. elegans longevity. Nat. Genet. 33, 40–48 (2003).
Liu, X. et al. Evolutionary conservation of the clk-1-dependent mechanism of longevity: loss of mclk1 increases cellular fitness and lifespan in mice. Genes Dev. 19, 2424–2434 (2005).
Lakowski, B. & Hekimi, S. Determination of life-span in Caenorhabditis elegans by four clock genes. Science 272, 1010–1013 (1996).
Copeland, J. M. et al. Extension of Drosophila life span by RNAi of the mitochondrial respiratory chain. Curr. Biol. 19, 1591–1598 (2009).
Owusu-Ansah, E., Song, W. & Perrimon, N. Muscle mitohormesis promotes longevity via systemic repression of insulin signaling. Cell 155, 699–712 (2013).
Delaney, J. R. et al. Stress profiling of longevity mutants identifies Afg3 as a mitochondrial determinant of cytoplasmic mRNA translation and aging. Aging Cell 12, 156–166 (2013).
Yee, C., Yang, W. & Hekimi, S. The intrinsic apoptosis pathway mediates the pro-longevity response to mitochondrial ROS in C. elegans. Cell 157, 897–909 (2014).
Labbadia, J. & Morimoto, R. I. Repression of the heat shock response is a programmed event at the onset of reproduction. Mol. Cell 59, 639–650 (2015).
Dell'agnello, C. et al. Increased longevity and refractoriness to Ca2+-dependent neurodegeneration in Surf1 knockout mice. Hum. Mol. Genet. 16, 431–444 (2007).
Munkacsy, E. & Rea, S. L. The paradox of mitochondrial dysfunction and extended longevity. Exp. Gerontol. 56, 221–233 (2014).
Ren, Y. et al. The activation of protein homeostasis protective mechanisms perhaps is not responsible for lifespan extension caused by deficiencies of mitochondrial proteins in C. elegans. Exp. Gerontol. 65, 53–57 (2015).
Schieber, M. & Chandel, N. S. TOR signaling couples oxygen sensing to lifespan in C. elegans. Cell Rep. 9, 9–15 (2014).
Bennett, C. F. et al. Transaldolase inhibition impairs mitochondrial respiration and induces a starvation-like longevity response in Caenorhabditis elegans. PLoS Genet. 13, e1006695 (2017).
Lee, S. J., Hwang, A. B. & Kenyon, C. Inhibition of respiration extends C. elegans life span via reactive oxygen species that increase HIF-1 activity. Curr. Biol. 20, 2131–2136 (2010).
Yun, J. & Finkel, T. Mitohormesis. Cell Metab. 19, 757–766 (2014).
Beck, J. S., Mufson, E. J. & Counts, S. E. Evidence for mitochondrial UPR gene activation in familial and sporadic Alzheimer's disease. Curr. Alzheimer Res. 13, 610–614 (2016).
Kambe, Y. & Miyata, A. Potential involvement of the mitochondrial unfolded protein response in depressive-like symptoms in mice. Neurosci. Lett. 588, 166–171 (2015).
Angelastro, J. M. Targeting ATF5 in Cancer. Trends Cancer 3, 471–474 (2017).
Mouchiroud, L. et al. The NAD+/sirtuin pathway modulates longevity through activation of mitochondrial UPR and FOXO signaling. Cell 154, 430–441 (2013).
Zhang, H. et al. NAD+ repletion improves mitochondrial and stem cell function and enhances life span in mice. Science 352, 1436–1443 (2016).
Gariani, K. et al. Eliciting the mitochondrial unfolded protein response by nicotinamide adenine dinucleotide repletion reverses fatty liver disease in mice. Hepatology 63, 1190–1204 (2016).
Tanaka, T. et al. Targeted disruption of ATF4 discloses its essential role in the formation of eye lens fibres. Genes Cells 3, 801–810 (1998).
Dalton, R. P., Lyons, D. B. & Lomvardas, S. Co-opting the unfolded protein response to elicit olfactory receptor feedback. Cell 155, 321–332 (2013).
Wang, S. Z., Ou, J., Zhu, L. J. & Green, M. R. Transcription factor ATF5 is required for terminal differentiation and survival of olfactory sensory neurons. Proc. Natl Acad. Sci. USA 109, 18589–18594 (2012).
West, A. P., Shadel, G. S. & Ghosh, S. Mitochondria in innate immune responses. Nat. Rev. Immunol. 11, 389–402 (2011).
Montecino-Rodriguez, E., Berent-Maoz, B. & Dorshkind, K. Causes, consequences, and reversal of immune system aging. J. Clin. Invest. 123, 958–965 (2013).
Rudel, T., Kepp, O. & Kozjak-Pavlovic, V. Interactions between bacterial pathogens and mitochondrial cell death pathways. Nat. Rev. Microbiol. 8, 693–705 (2010).
West, A. P. & Shadel, G. S. Mitochondrial DNA in innate immune responses and inflammatory pathology. Nat. Rev. Immunol. 17, 363–375 (2017).
Galmiche, A. & Rassow, J. Targeting of Helicobacter pylori VacA to mitochondria. Gut Microbes 1, 392–395 (2010).
Bos, L. D., Sterk, P. J. & Schultz, M. J. Volatile metabolites of pathogens: a systematic review. PLoS Pathog. 9, e1003311 (2013).
Melo, J. A. & Ruvkun, G. Inactivation of conserved C. elegans genes engages pathogen- and xenobiotic-associated defenses. Cell 149, 452–466 (2012).
Reddy, K. C., Dunbar, T. L., Nargund, A. M., Haynes, C. M. & Troemel, E. R. The C. elegans CCAAT-enhancer-binding protein gamma is required for surveillance immunity. Cell Rep. 14, 1581–1589 (2016).
Rehacek, Z., Ramankutty, M. & Kozova, J. Respiratory chain of antimycin A-producing Streptomyces antibioticus. Appl. Microbiol. 16, 29–32 (1968).
Smith, R. M., Peterson, W. H. & McCoy, E. Oligomycin, a new antifungal antibiotic. Antibiot. Chemother. 4, 962–970 (1954).
Denver, D. R., Morris, K., Lynch, M., Vassilieva, L. L. & Thomas, W. K. High direct estimate of the mutation rate in the mitochondrial genome of Caenorhabditis elegans. Science 289, 2342–2344 (2000).
Stewart, J. B. & Chinnery, P. F. The dynamics of mitochondrial DNA heteroplasmy: implications for human health and disease. Nat. Rev. Genet. 16, 530–542 (2015).
Payne, B. A. et al. Universal heteroplasmy of human mitochondrial DNA. Hum. Mol. Genet. 22, 384–390 (2013).
Kauppila, T. E., Kauppila, J. H. & Larsson, N. G. Mammalian mitochondria and aging: an update. Cell Metab. 25, 57–71 (2017).
Tsang, W. Y. & Lemire, B. D. Stable heteroplasmy but differential inheritance of a large mitochondrial DNA deletion in nematodes. Biochem. Cell Biol. 80, 645–654 (2002).
Pulliam, D. A. et al. Complex IV-deficient Surf1(−/−) mice initiate mitochondrial stress responses. Biochem. J. 462, 359–371 (2014).
Khan, N. A. et al. Effective treatment of mitochondrial myopathy by nicotinamide riboside, a vitamin B3 . EMBO Mol. Med. 6, 721–731 (2014).
Rainbolt, T. K., Atanassova, N., Genereux, J. C. & Wiseman, R. L. Stress-regulated translational attenuation adapts mitochondrial protein import through Tim17A degradation. Cell Metab. 18, 908–919 (2013).
Acknowledgements
This work was supported by an EMBO long-term fellowship (ALTF 715–2015) to T.S., the Mallinckrodt Foundation, HHMI and NIH grants R01AG040061, R01AG047182 and R01HL127891 to C.M.H.
Author information
Authors and Affiliations
Contributions
T.S. and C.M.H. researched data for the article, contributed to discussion of the content, wrote the article and reviewed and/or edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Oxidative phosphorylation
-
(OXPHOS). The mitochondrial pathway that metabolizes nutrients and generates ATP, which requires the respiratory chain and ATP synthase complexes.
- Translocase of the outer membrane complex
-
(TOM complex). A protein complex localized in the mitochondrial outer membrane required for nuclear-encoded proteins synthesized on cytosolic ribosomes to cross the mitochondrial outer membrane.
- Translocase of the inner membrane 23 complex
-
(TIM23 complex). A protein complex localized in the mitochondrial inner membrane that facilitates the translocation of proteins from the intermembrane space to the mitochondrial matrix or to the mitochondrial inner membrane.
- Integrated stress response
-
(ISR). A stress response initiated by kinases responsive to endoplasmic reticulum stress, amino acid depletion, haem depletion or viral infection that leads to phosphorylation of eukaryotic translation initiation factor 2 subunit 1 (eIF2α), ultimately resulting in reduced protein synthesis and increased translation of mRNAs harbouring upstream open reading frames.
- β-Oxidation
-
The catabolic process that occurs within mitochondria by which the breakdown of fatty acids yields acetyl-CoA, NADH and FADH2.
- Adipose tissue browning
-
The development of beige adipocytes in white adipose tissue, which involves the accumulation of mitochondria within white adipocytes.
- Hypoxia-inducible factor 1
-
(HIF-1). A transcription factor activated during hypoxia that promotes cell growth and survival by affecting a variety of processes, including metabolic adaptations.
- Heteroplasmy
-
The presence of more than one type of mitochondrial DNA within a cell or individual.
Rights and permissions
About this article
Cite this article
Shpilka, T., Haynes, C. The mitochondrial UPR: mechanisms, physiological functions and implications in ageing. Nat Rev Mol Cell Biol 19, 109–120 (2018). https://doi.org/10.1038/nrm.2017.110
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm.2017.110
This article is cited by
-
Mitochondrial stress activates YAP/TAZ through RhoA oxidation to promote liver injury
Cell Death & Disease (2024)
-
Bacterial peptidoglycan acts as a digestive signal mediating host adaptation to diverse food resources in C. elegans
Nature Communications (2024)
-
The mitochondrial UPR induced by ATF5 attenuates intervertebral disc degeneration via cooperating with mitophagy
Cell Biology and Toxicology (2024)
-
(+)-Lipoic acid reduces mitochondrial unfolded protein response and attenuates oxidative stress and aging in an in vitro model of non-alcoholic fatty liver disease
Journal of Translational Medicine (2024)
-
ATF5 regulates tubulointerstitial injury in diabetic kidney disease via mitochondrial unfolded protein response
Molecular Medicine (2023)